Controlling the environment of an application is vital for it’s functionality and stability. Especially in a distributed environment it is important for developers to have control over the version of dependencies. In such an scenario it’s a critical task to ensure possible conflicting requirements of multiple applications are not disturbing each other.
That is why frameworks like YARN ensure that each application is executed in a self-contained environment – typically in a Linux Container or Docker Container – that is controlled by the developer. In this post we show what this means for Python environments being used by Spark.
YARN Application Deployment
As mentioned earlier does YARN execute each application in a self-contained environment on each host. This ensures the execution in a controlled environment managed by individual developers. The way this works in a nutshell is that the dependency of an application are distributed to each node typically via HDFS.
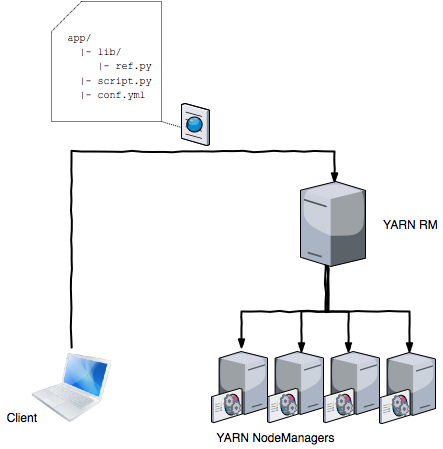
The files are uploaded to a staging folder /user/${username}/.${application} of the submitting user in HDFS. Because of the distributed architecture of HDFS it is ensured that multiple nodes have local copies of the files. In fact to ensure that a large fraction of the cluster has a local copy of application files and does not need to download them over the network, the HDFS replication factor is set much higher for this files than 3. Often a number between 10 and 20 is chosen for the replication factor.
During the preparation of the container on a node you will notice in logs similar commands to the below example are being executed:
ln -sf "/hadoop/yarn/local/usercache/vagrant/filecache/72/pyspark.zip" "pyspark.zip"
The folder /hadoop/yarn/local/ is the configured location on each node where YARN stores it’s needed files and logs locally. Creating a symbolic link like this inside the container makes the content of the zip file available. It is being referenced as “pyspark.zip”.
Using Conda Env
For application developers this means that they can package and ship their controlled environment with each application. Other solutions like NFS or Amazon EFS shares are not needed, especially since solutions like shared folders makes for a bad architecture that is not designed to scale very well making the development of application less agile.
The following example demonstrate the use of conda env to transport a python environment with a PySpark application needed to be executed. This sample application uses the NLTK package with the additional requirement of making tokenizer and tagger resources available to the application as well.
Our sample application:
import os
import sys
from pyspark import SparkContext
from pyspark import SparkConf
conf = SparkConf()
conf.setAppName("spark-ntlk-env")
sc = SparkContext(conf=conf)
data = sc.textFile('hdfs:///user/vagrant/1970-Nixon.txt')
def word_tokenize(x):
import nltk
return nltk.word_tokenize(x)
def pos_tag(x):
import nltk
return nltk.pos_tag([x])
words = data.flatMap(word_tokenize)
words.saveAsTextFile('hdfs:///user/vagrant/nixon_tokens')
pos_word = words.map(pos_tag)
pos_word.saveAsTextFile('hdfs:///user/vagrant/nixon_token_pos')
Preparing the sample input data
For our example we are using the provided samples of NLTK (http://www.nltk.org/nltk_data/) and upload them to HDFS:
(nltk_env)$ python -m nltk.downloader -d nltk_data all (nltk_env)$ hdfs dfs -put nltk_data/corpora/state_union/1970-Nixon.txt /user/vagrant/
No Hard (Absolute) Links!
Before we actually go and create our environment lets first take a quick moment to recap on how an environment is typically being composed. On a machine the environment is made out of variables linking to different target folders containing executable or other resource files. So if you execute a command it is either referenced from your PATH, PYTHON_LIBRARY, or any other defined variable. These variables link to files in directories like /usr/bin, /usr/local/bin or any other referenced location. They are called hard links or absolute reference as they start from root /.
Environments using hard links are not easily transportable as they make strict assumption about the the overall execution engine (your OS for example) they are being used in. Therefor it is necessary to use relative links in a transportable/relocatable environment.
This is especially true for conda env as it creates hard links by default. By making the conda env relocatable it can be used in a application by referencing it from the application root . (current dir) instead of the overall root /. By using the –copy options during the creation of the environment packages are being copied instead of being linked.
Creating our relocatable environment together with nltk and numpy:
conda create -n nltk_env --copy -y -q python=3 nltk numpy
Fetching package metadata .......
Solving package specifications: ..........
Package plan for installation in environment /home/datalab_user01/anaconda2/envs/nltk_env:
The following packages will be downloaded:
package | build
---------------------------|-----------------
python-3.5.2 | 0 17.2 MB
nltk-3.2.1 | py35_0 1.8 MB
numpy-1.11.1 | py35_0 6.1 MB
setuptools-23.0.0 | py35_0 460 KB
wheel-0.29.0 | py35_0 82 KB
pip-8.1.2 | py35_0 1.6 MB
------------------------------------------------------------
Total: 27.2 MB
The following NEW packages will be INSTALLED:
mkl: 11.3.3-0 (copy)
nltk: 3.2.1-py35_0 (copy)
numpy: 1.11.1-py35_0 (copy)
openssl: 1.0.2h-1 (copy)
pip: 8.1.2-py35_0 (copy)
python: 3.5.2-0 (copy)
readline: 6.2-2 (copy)
setuptools: 23.0.0-py35_0 (copy)
sqlite: 3.13.0-0 (copy)
tk: 8.5.18-0 (copy)
wheel: 0.29.0-py35_0 (copy)
xz: 5.2.2-0 (copy)
zlib: 1.2.8-3 (copy)
#
# To activate this environment, use:
# $ source activate nltk_env
#
# To deactivate this environment, use:
# $ source deactivate
#
This also works for different python version 3.x or 2.x!
Zip it and Ship it!
Now that we have our relocatable environment all set we are able to package it and ship it as part of our sample PySpark job.
$ cd ~/anaconda2/envs/ $ zip -r nltk_env.zip nltk_env
Making this available in during the execution of our application in a YARN container we have for one distribute the package and for second change the default environment of Spark for python to your location.
The variable controlling the python environment for python applications in Spark is named PYSPARK_PYTHON.
PYSPARK_PYTHON=./NLTK/nltk_env/bin/python spark-submit --conf spark.yarn.appMasterEnv.PYSPARK_PYTHON=./NLTK/nltk_env/bin/python --master yarn-cluster --archives nltk_env.zip#NLTK spark_nltk_sample.py
Our virtual environment is linked by NLTK that is why the path in PYSPARK_PYTHON is pointing to ./NLTK/content/of/zip/… . The exact command being executed during container creation is something like this:
ln -sf "/hadoop/yarn/local/usercache/vagrant/filecache/71/nltk_env.zip" "NLTK"
Shipping additional resources with an application is controlled by the –files and –archives options as shown here.
The options being used here are documented in Spark Yarn Configuration and Spark Environment Variables for reference.
Packaging tokenizer and taggers
Doing just the above will unfortunately fail, because using the NLTK parser in the way we are using it in the example program has some additional dependencies. If you have followed the above steps executing submitting it to your YARN cluster will result in the following exception at container level:
Caused by: org.apache.spark.api.python.PythonException: Traceback (most recent call last):
...
raise LookupError(resource_not_found)
LookupError:
**********************************************************************
Resource 'taggers/averaged_perceptron_tagger/averaged_perceptron
_tagger.pickle' not found. Please use the NLTK Downloader to
obtain the resource: >>> nltk.download()
Searched in:
- '/home/nltk_data'
- '/usr/share/nltk_data'
- '/usr/local/share/nltk_data'
- '/usr/lib/nltk_data'
- '/usr/local/lib/nltk_data'
**********************************************************************
at org.apache.spark.api.python.PythonRunner$$anon$1.read(PythonRDD.scala:166)
at org.apache.spark.api.python.PythonRunner$$anon$1.<init>(PythonRDD.scala:207)
at org.apache.spark.api.python.PythonRunner.compute(PythonRDD.scala:125)
at org.apache.spark.api.python.PythonRDD.compute(PythonRDD.scala:70)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:300)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:264)
at org.apache.spark.rdd.MapPartitionsRDD.compute(MapPartitionsRDD.scala:38)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:300)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:264)
at org.apache.spark.rdd.MapPartitionsRDD.compute(MapPartitionsRDD.scala:38)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:300)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:264)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:66)
at org.apache.spark.scheduler.Task.run(Task.scala:88)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:214)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1142)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:617)
... 1 more
The problem is that NLTK expects the follwoing resource in the tokenizers/punkt/english.pickle to be available in either of the following locations:
Searched in:
- '/home/nltk_data'
- '/usr/share/nltk_data'
- '/usr/local/share/nltk_data'
- '/usr/lib/nltk_data'
- '/usr/local/lib/nltk_data'
The good thing about this is, that we by now should now how we can ship the required dependency to our application. We can do it the same way we did with our python environment.
Again it is imporant to reensure yourself how the resource is going to be referenced. NLTK expects it by default in the current location under tokenizers/punkt/english.pickle that is why we navigate into the folder for packaging and reference the zip file wiht tokenizer.zip#tokenizer.
(nltk_env)$ cd nltk_data/tokenizers/ (nltk_env)$ zip -r ../../tokenizers.zip * (nltk_env)$ cd ../../ (nltk_env)$ cd nltk_data/taggers/ (nltk_env)$ zip -r ../../taggers.zip * (nltk_env)$ cd ../../
At a later point our program will expect a tagger in the same fashion already demonstrated in the above snippet.
Using YARN Locations
We can ship those zip resources the same way we shipped our conda env. In addition environment variable can be used to control resource discovery and allocation.
For NLTK you can use the environment variable NLTK_DATA to control the path. Setting this in Spark can be done similar to the way we set PYSPARK_PYTHON:
--conf spark.yarn.appMasterEnv.NLTK_DATA=./
Additionally YARN exposes the container path via the environment variable PWD. This can be used in NLTK as to add it to the search path as follows:
def word_tokenize(x):
import nltk
nltk.data.path.append(os.environ.get('PWD'))
return nltk.word_tokenize(x)
The submission of your application becomes now:
PYSPARK_PYTHON=./NLTK/nltk_env/bin/python spark-submit --conf spark.yarn.appMasterEnv.PYSPARK_PYTHON=./NLTK/nltk_env/bin/python --conf spark.yarn.appMasterEnv.NLTK_DATA=./ --master yarn-cluster --archives nltk_env.zip#NLTK,tokenizers.zip#tokenizers,taggers.zip#taggers spark_nltk_sample.py
The expected result should be something like the following:
(nltk_env)$ hdfs dfs -cat /user/datalab_user01/nixon_tokens/* | head -n 20 Annual Message to the Congress on the State of the Union . January 22 , 1970 Mr. Speaker , Mr.
And:
(nltk_env)$ hdfs dfs -cat /user/datalab_user01/nixon_token_pos/* | head -n 20 [(u'Annual', 'JJ')] [(u'Message', 'NN')] [(u'to', 'TO')] [(u'the', 'DT')] [(u'Congress', 'NNP')] [(u'on', 'IN')] [(u'the', 'DT')] [(u'State', 'NNP')] [(u'of', 'IN')] [(u'the', 'DT')] [(u'Union', 'NN')] [(u'.', '.')] [(u'January', 'NNP')] [(u'22', 'CD')] [(u',', ',')] [(u'1970', 'CD')] [(u'Mr.', 'NNP')] [(u'Speaker', 'NN')] [(u',', ',')] [(u'Mr.', 'NNP')]
3 thoughts on “Running PySpark with Conda Env”